Attempting to debug dynlib loading issues in Windows can be tricky – on Linux you can use strace to monitor dynamic linking and see what libraries are being loaded as the application runs. Having just had some issues with dynlib loading issues in an R extension I wrote, I was keen to find a good way to debugging the issues as they occurred. Thankfully, there are some good utilities out there for debugging this kind of thing:
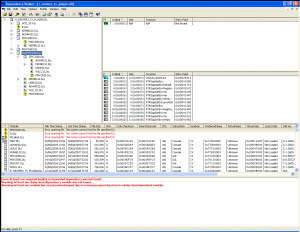
Dependency Walker is an old SDK tool that is still invaluable. Drag a .DLL or .EXE into the application window and it will show you the dependencies, what type of loading mechanism they use (e.g. eager or delay load), and also what exported functions from the dependent DLLs are actually used by the .DLL or .EXE you have selected.
Here is a screenshot of depends.exe operating on my R extension DLL. Note that MSCVR80.DLL and R.DLL are both highlighted as missing – this is extremely useful for figuring out dependent DLL load issues.
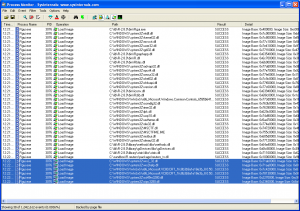
The SysInternals Process Monitor is an incredibly useful low-level Swiss Army Knife utility that can be used, among other things, to monitor dynamic library loading activity as it occurs, using the file activity view. Here is a view of Process Monitor monitoring R as it loads my extension DLL. The subsequent dependent DLL loading activity is highlighted.